Shelby Whitaker ’24: The Burns Lab at Boston Children’s Hospital and Harvard Medical School
- The Rivers School

- Aug 18, 2023
- 4 min read
Growing up, I have been a frequent flier at Children's Hospital for my many sports-related injuries. I have always been fascinated with how hospitals work after having a first-class seat to observe the miracles of medicine. Behind the scenes, there are many incredible labs working on the next medical discovery to revolutionize new patient care. So, I was very excited to have the opportunity to intern at the Burns Lab at Boston Children's Hospital this summer. The Burns Lab is a cardiology lab that uses zebrafish as a model organism to study human cardiovascular diseases. Zebrafish are an ideal model because they have a similar genetic structure to humans and they are highly regenerative.
During my internship, I worked with postdoctoral fellow Hakan Coskun to characterize a new zebrafish model of Williams Syndrome. Williams Syndrome is caused by a missing segment or deletion of genetic material on chromosome 7. This deletion involves 26–28 genes, including the Elastin (ELN) gene. This gene contributes to devastating cardiovascular complications in patients with this syndrome. The goal of my project was to analyze the cardiovascular phenotypes of zebrafish embryos missing the equivalent of the human ELN gene.
Where all the zebrafish are kept
Creating the genetically modified zebrafish is a challenge because, within the elastin gene, there are two isoforms, ElnA and ElnB. Each isoform has two copies. Therefore, to have a true loss-of-function fish, all 4 alleles must be knocked out. Hakan previously used CRISPR-Cas9 genome editing to create the strain of mutated zebrafish. In order to check if the mutation was successful I genotyped the animals. The first step is to obtain DNA from the fish by cutting a little piece of the fin. Next, I ran a PCR (polymerase chain reaction). PCR is a technique to rapidly produce (amplifying) millions to billions of copies of a specific segment of DNA, which can then be studied in greater detail. Dr. Coskun took the time to teach me the specifics of this process which involves three steps, including denaturation, annealing, and extension. Each step occurs at a set temperature for different durations of time in a PCR machine.
L: Loading the DNA samples into the gel; R: Picture of one of the PCR results
After the PCR was complete I would run these DNA samples through gel electrophoresis. This process would allow me to visualize the DNA fragment and its size. As a result, I found many fish that were ElnA +/- ElnB -/- . This means that these fish had no copies of the Elastin B gene and only one out of the two copies of Elastin A. I then set up an incross between these adult fish and collected their embryos. The great thing about this cross is that we are ensured to have no ElnB so we have greater chances to get double mutants.
These embryos also contained a fluorescent transgene called fggr. Fggr makes it so that when viewing the embryos under a microscope, I was able to clearly see green fluorescence marking the blood vessel cells (endothelium) and red fluorescence in the circulating blood cells. When the embryos were seven days post fertilization (7dpf), I took pictures and videos of these fish using the microscope to then be later analyzed. Once that was finished I ran yet another PCR using the gel to figure out what genotype they were.
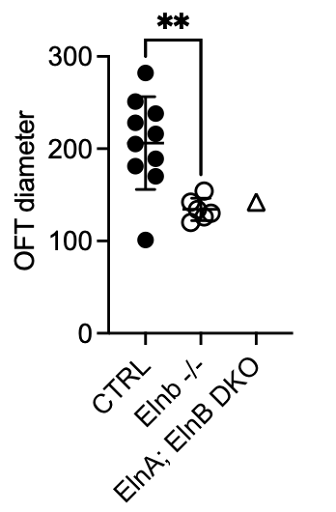
Putting it all together I used the videos to compare the mutant zebrafish with the wild type (ElnA +/+ and ElnB +/+). Specifically, I quantified the differences in the diameter of the outflow tract, which is equivalent to the base of the aorta in humans.
L: Imaging my embryos; R: Results of my measurements
From my work, I discovered that only Elastin B affects outflow tract development. With the absence of Elastin B, the smooth muscle cells do not form, which leads to a rigid structure and reduced blood circulation.
In addition to working in the lab, all of the summer interns working in the Department of Cardiology met with Dr. Vassilios Bezzerides once a week for journal club. Each week we would read an original scientific research paper and would be assigned a specific figure to present to the entire group. These papers were very difficult to understand as they were many pages long filled with graphs, images, and statistics. As the weeks progressed, I learned what I needed to focus on in my section to be able to glean the important information.
Through journal club, I was also fortunate enough to be able to shadow Dr. Bezzerides to see the clinical side of his work. I was able to see him perform a procedure in an electrophysiology (EP) lab where he induced a patient's irregular arrhythmia so he could use ablations to fix the heart. After that, he showed me around different operating rooms (ORs) and I even saw a small part of an open heart surgery on a two-year-old patient. I was able to appreciate the numerous and diverse medical professionals and the significant logistical choreography at work during each procedure.
Throughout my seven weeks in the Burns Lab, I learned so much about research. More specifically I learned many technical skills like using lab equipment, running experiments, and interpreting results. Additionally, when putting my results together I learned about the importance of quantifying your data and how best to represent it.
I am very grateful to have been afforded the opportunity to learn about many different career paths from some very kind and intelligent individuals. I am beyond appreciative that so many gifted and talented professionals were willing to share their knowledge with me. I would like to thank everyone in the lab, especially Dr. Caroline Burns and Dr. Geoff Burns, who welcomed me into their lab and willingly jumped in and helped me whenever I didn't know what to do (which was often!). And a special thanks to Dr. Hakan Coskun who made my experience so unique. This internship has given me so much experience and information that will only help me grow my interest in the STEM field. Thank you also to Mr. Schlenker for providing me with this amazing opportunity.
















Comments